The complex genetic architecture of adaptive traits
What limits the speed of adaptation in a changing world? A key constraint is the underlying genetic architecture – the number of required genetic changes and their individual effects across environments. These architectures are influenced by variation at temporal and spatial scales (check out my new perspective piece in New Phyt for more!). In the Kreiner lab, we quantify adaptive genetic architectures and their variability to better understand key determinants and predict the pace of adaptation across heterogeneous landscapes.
Herbicide resistance
Mutations at genes that code for enzymes targeted by herbicides have by and large been the major focus of the study of herbicide resistance in weed populations. This tends to be because these causal mutations are easy to assay. For example, in our work on the adaptive origins of the amplification conferring glyphosate resistance (described above), I was able to focus my investigations to a very particular region of the genome. However, there are numerous other physiological mechanisms that confer resistance in natural populations—mechanisms for which we have no idea what loci and genes underlie this response. In our study populations of common waterhemp, a substantial portion of the variation in herbicide resistance is unexplained by known target-site mechanisms. In the map above, I’ve illustrated not only this point – that many individuals show resistance but lack known target-site mechanism -and therefor the extent that the genetic architecture of resistance varies across populations.
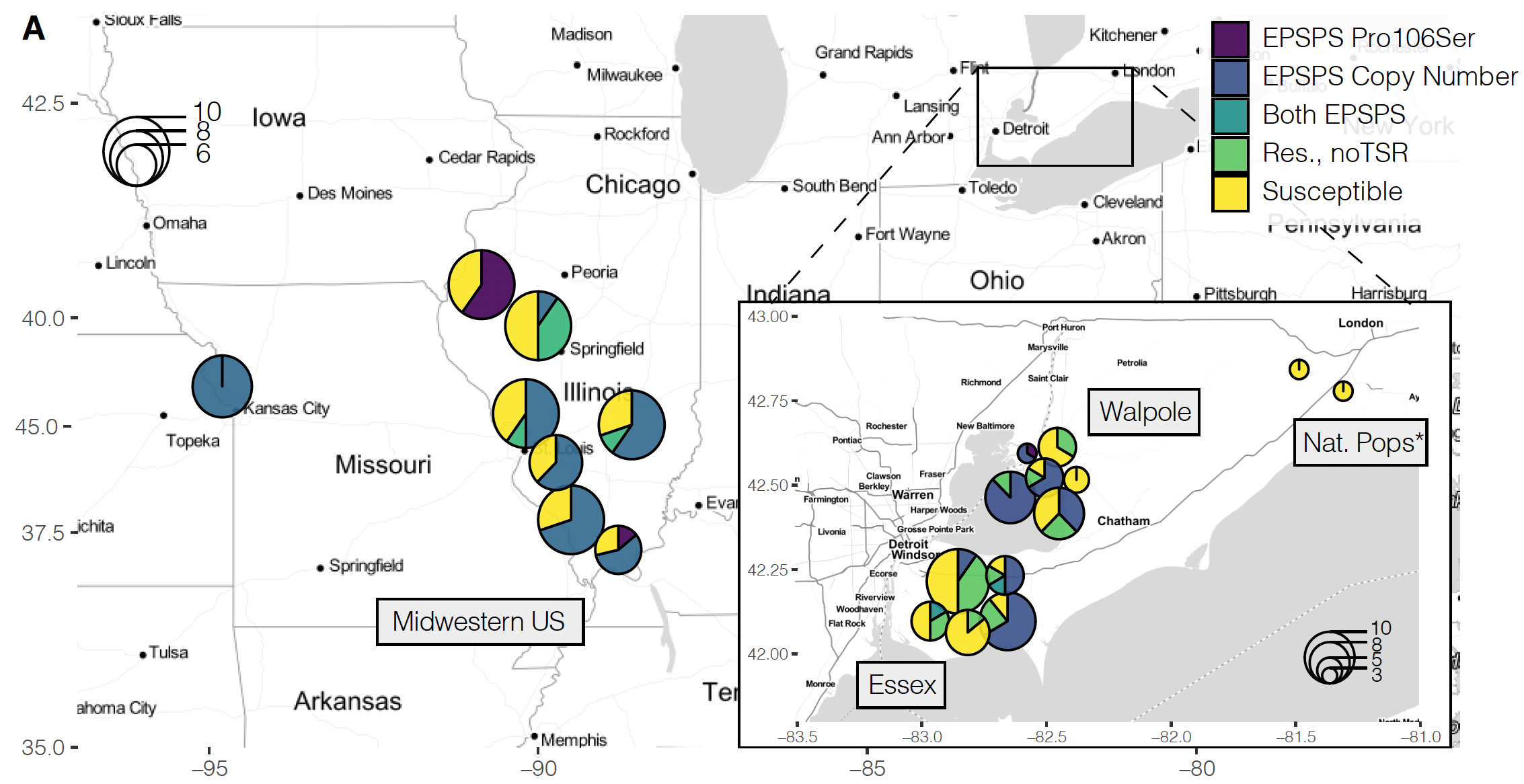
We therefore set out to characterize the polygenic basis of resistance in common waterhemp, both with regard to genetic architecture of herbicide resistance (the number and distribution of genes involved, the frequency and effect size of resistance-loci) and how selection from herbicides has shaped genome-wide diversity. In particular, I was interested in how much variation in resistance across populations could be explained by genome-wide loci compared to monogenic mechanisms that are typically the focus of herbicide resistance research. In our paper at Molecular Ecology, published as a part of the special issue “Resistance evolution, from molecular mechanism to ecological context”, curated by Gina Baucom], we quantify the relative importance of polygenic vs monogenic herbicide resistance, show that strong selection shapes its genetic architecture with large effect & rare alleles, and find evidence of their maintenance by recent positive selection. Somewhat surprisingly, we show that polygenic resistance is of comparable importance to monogenic mechanisms in conferring round-up resistance in agricultural populations, even in individuals that already segregate for large-effect target-site mutations.
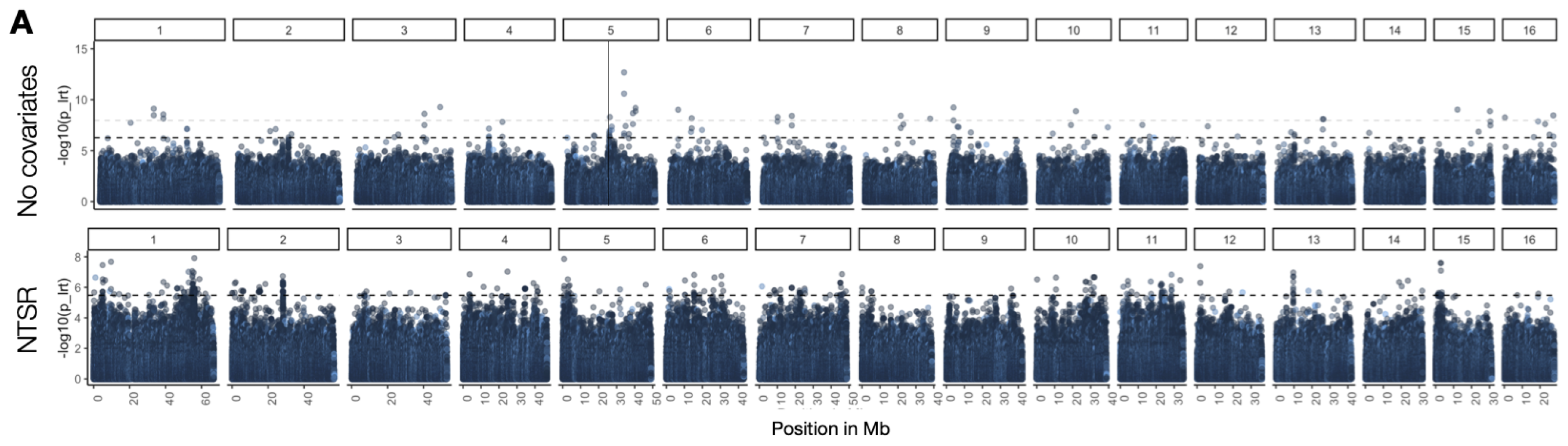
Flowering Time
Flowering time is one of the most important traits for determining fitness in plants. If you flower to early or late in the season, your most important asset will be at risk to the elements (frost!). The genetics of flowering time is pretty well characterized in some model species (Arabidopsis thaliana, the fruit fly of plants), but we had no idea about what determined variation in flowering time at the genetic level in Amaranthus tubercultus. While perhaps less pressing than understanding than herbicide resistance in terms of controlling them in problematic settings, we previously found that waterhemp has had to shift its flowering time in order to be well suited to grow in agricultural settings, meaning its also key to the bigger picture of management.
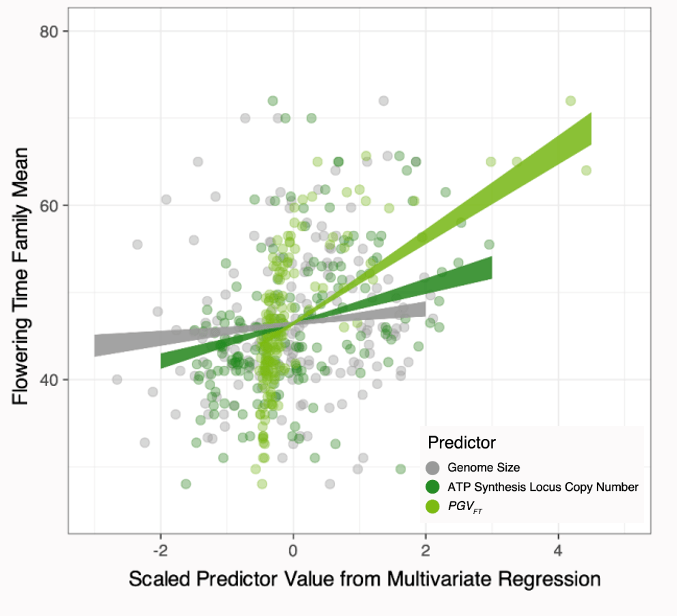
We took a multi-level approach to characterizing the genetics of flowering time in Amaranthus. We implemented GWAS for SNPs, copy number variants, and looked at the quantative effect of genome size on flowering time to describe a complex architure that draws on multiple types of genetic variation and is influenced by numerous processes across the genome and across the geographic range.
For more details, please check out or recent preprint!
A. tuberculatus
Relevant publications
Kreiner JM. 2025. The genetic architecture and spatiotemporal dynamics of adaptation across human-modified landscapes. New Phytologist
Kreiner JM, Tranel PJ, Weigel D, Stinchcombe JR, & Wright SI. 2021. The genetic architecture and population genomic signatures of glyphosate resistance. Molecular Ecology
Kreiner JM, Hnatovska S, Stinchcombe JR, & Wright SI. Genome size and repeat content contribute to a complex architecture of flowering time in Amaranthus tuberculatus. PLOS Genetics
